 |
biomcmc-lib
0.1
low level library for phylogenetic analysis
|
 |
biomcmc-lib
0.1
low level library for phylogenetic analysis
|
branch length operations on topologies, including patristic distances More...
#include "topology_common.h"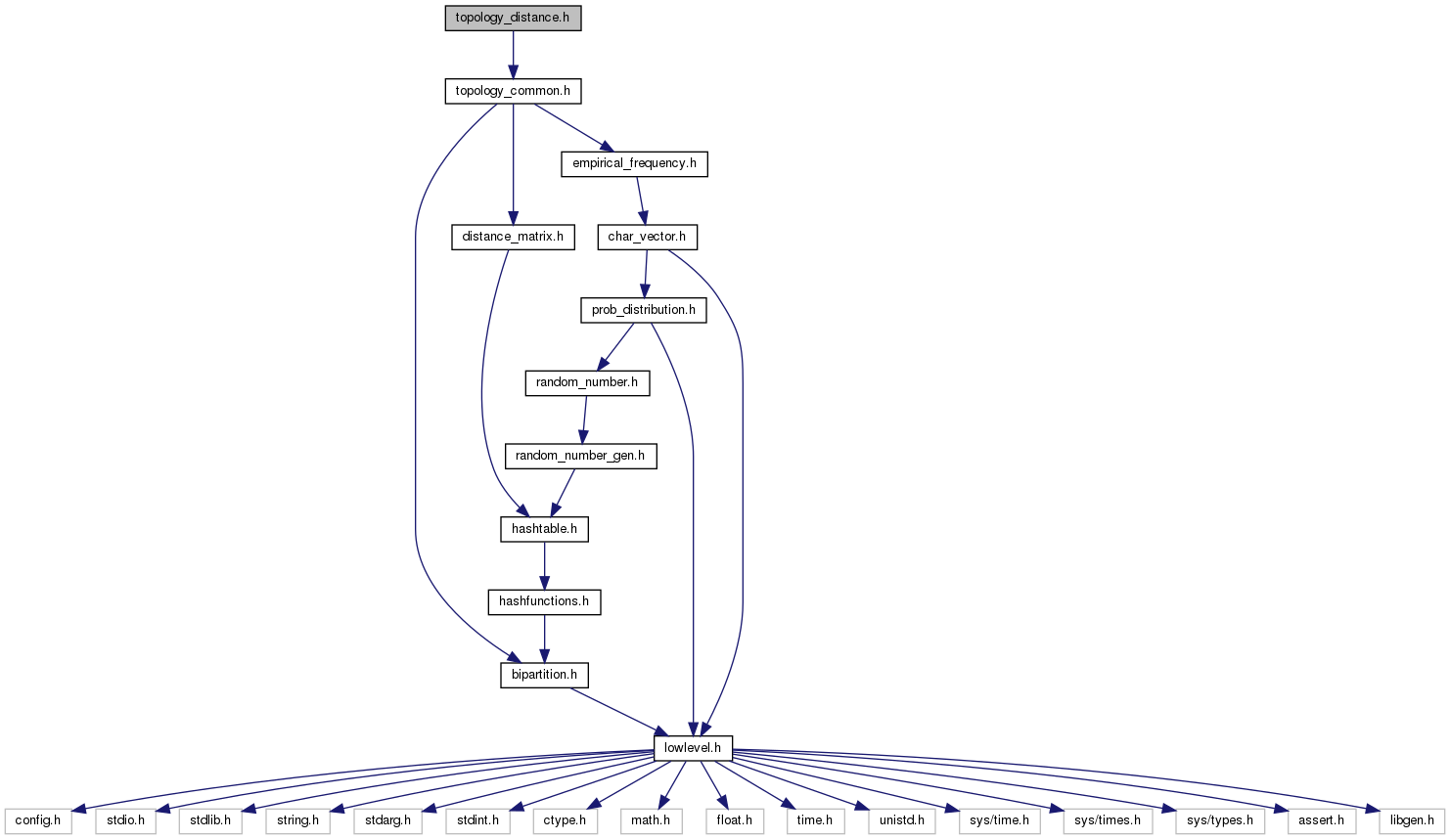
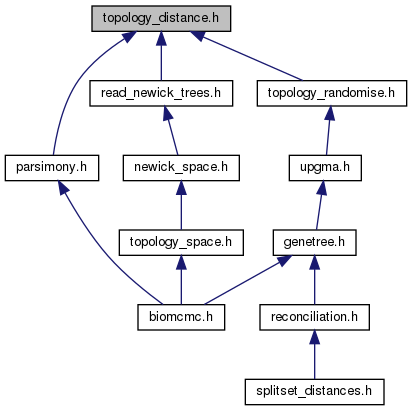
Go to the source code of this file.
Functions | |
| distance_matrix | new_distance_matrix_for_topology (int nleaves) |
| allocate memory for a new distance_matrix that will be used on topologies | |
| void | fill_distance_matrix_from_topology (distance_matrix dist, topology tree, double *blen, bool use_upper) |
| fill in distance_matrix with the patristic distances from topology (can be used with distinct branch length vectors to fill upper and lower diagonals | |
| void | patristic_distances_from_topology_to_vectors (topology tree, double **dist, double *scaling, int n_dists, double tolerance) |
| calculates rescaled patristic distances returning up to 6 distinct 1D vectors #dist (externally allocated) The 'tolerance' is the minimum branch length to be considered a multifurcation (length zero) | |
| int * | create_vector_with_idx_leaves_below_for_patristic (topology tree) |
| similar to an Euler tour, has list of leaves below each node | |
| void | estimate_topology_branch_lengths_from_distances (topology tree, double *dist) |
| double * | new_topology_branch_lengths_from_distances (topology tree, double *dist) |
| void | correct_negative_branch_lengths_from_topology (topology t, double *blength) |
branch length operations on topologies, including patristic distances
 1.8.13
1.8.13