 |
biomcmc-lib
0.1
low level library for phylogenetic analysis
|
 |
biomcmc-lib
0.1
low level library for phylogenetic analysis
|
distance matrix, that can be used in alignments and trees, and patristic-distance based species distances More...
#include "hashtable.h"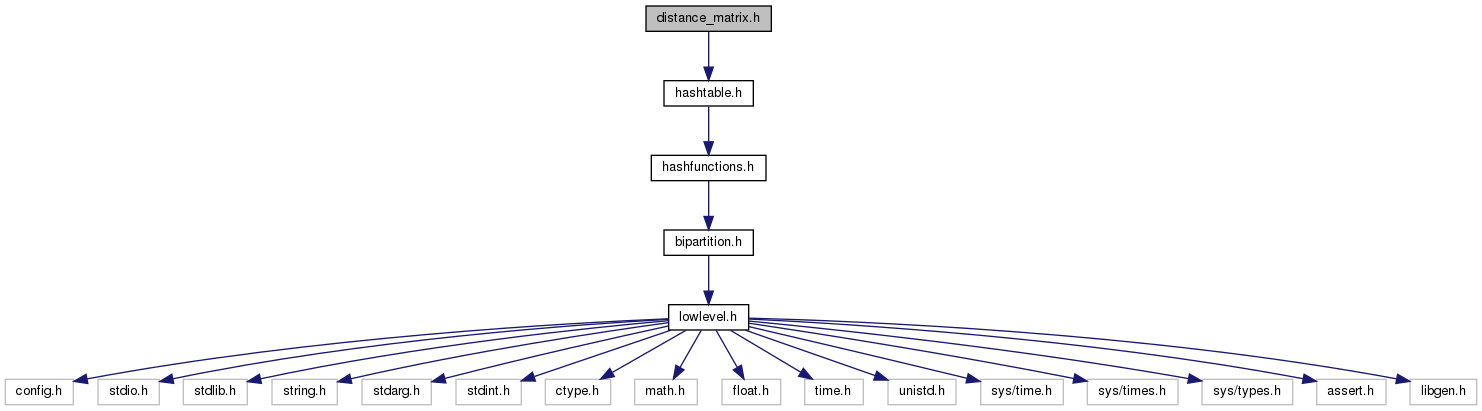
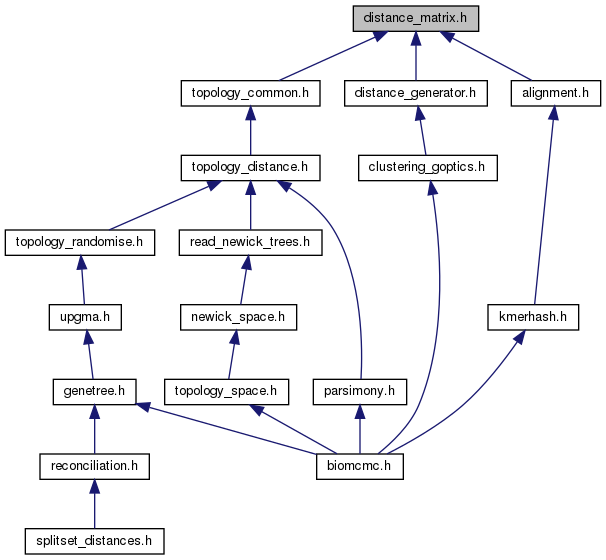
Go to the source code of this file.
Data Structures | |
| struct | distance_matrix_struct |
| struct | spdist_matrix_struct |
Typedefs | |
| typedef struct distance_matrix_struct * | distance_matrix |
| typedef struct spdist_matrix_struct * | spdist_matrix |
Functions | |
| distance_matrix | new_distance_matrix (int nseqs) |
| creates new matrix of pairwise distances | |
| void | zero_lower_distance_matrix (distance_matrix dist) |
| specially in gene/sptree distance methods (GLASS, STEAC, etc.) lower is used for means and upper for min. This function resets matrix elements | |
| void | transpose_distance_matrix (distance_matrix dist) |
| invert lower and upper diagonals of matrix (since some functions like upgma expect upper, etc.) | |
| void | del_distance_matrix (distance_matrix dist) |
| releases memory allocated to distance_matrix (this structure has no smart ref_counter) | |
| spdist_matrix | new_spdist_matrix (int n_species) |
| void | zero_all_spdist_matrix (spdist_matrix dist) |
| void | finalise_spdist_matrix (spdist_matrix dist) |
| void | finalise_spdist_matrix_with_rescaling (spdist_matrix dist, double scale) |
| void | complete_missing_spdist_from_global_spdist (spdist_matrix local, spdist_matrix global) |
| void | copy_spdist_matrix_to_distance_matrix_upper (spdist_matrix spd, distance_matrix dist, bool use_means) |
| void | del_spdist_matrix (spdist_matrix dist) |
| void | fill_species_dists_from_gene_dists (distance_matrix spdist, distance_matrix gendist, int *sp_id, bool use_upper_gene) |
| updates distances between species based on genes and gene-to-species mapping, with min on upper and mean on lower diagonal | |
| void | update_species_dists_from_spdist (distance_matrix global, distance_matrix local, int *spexist) |
| update global (over loci) species distances besed on local (within locus) species distances | |
| int | prepare_spdistmatrix_from_gene_species_map (spdist_matrix spdist, int *sp_id, int n_sp_id) |
| void | fill_spdistmatrix_from_gene_dists (spdist_matrix spdist, distance_matrix gendist, int *sp_id, bool use_upper_gene) |
| void | fill_spdistmatrix_from_gene_dist_vector (spdist_matrix spdist, double *gdist, int n_gdist, int *sp_id) |
| initialise spdist_matrix with patristic distances from gdist vector of size n_gdist (1D) | |
| void | update_spdistmatrix_from_spdistmatrix (spdist_matrix global, spdist_matrix local) |
distance matrix, that can be used in alignments and trees, and patristic-distance based species distances
These functions don't know about trees/topologies: topology_common.c creates the actual patristic distances, and downstream software or genetree.h should decide how to use this information
| void zero_all_spdist_matrix | ( | spdist_matrix | dist | ) |
zero both mean[] and min[] since we only look at average (never min) across loci
 1.8.13
1.8.13