 |
biomcmc-lib
0.1
low level library for phylogenetic analysis
|
 |
biomcmc-lib
0.1
low level library for phylogenetic analysis
|
vector of strings (species names, leaf names, etc.) More...
#include "char_vector.h"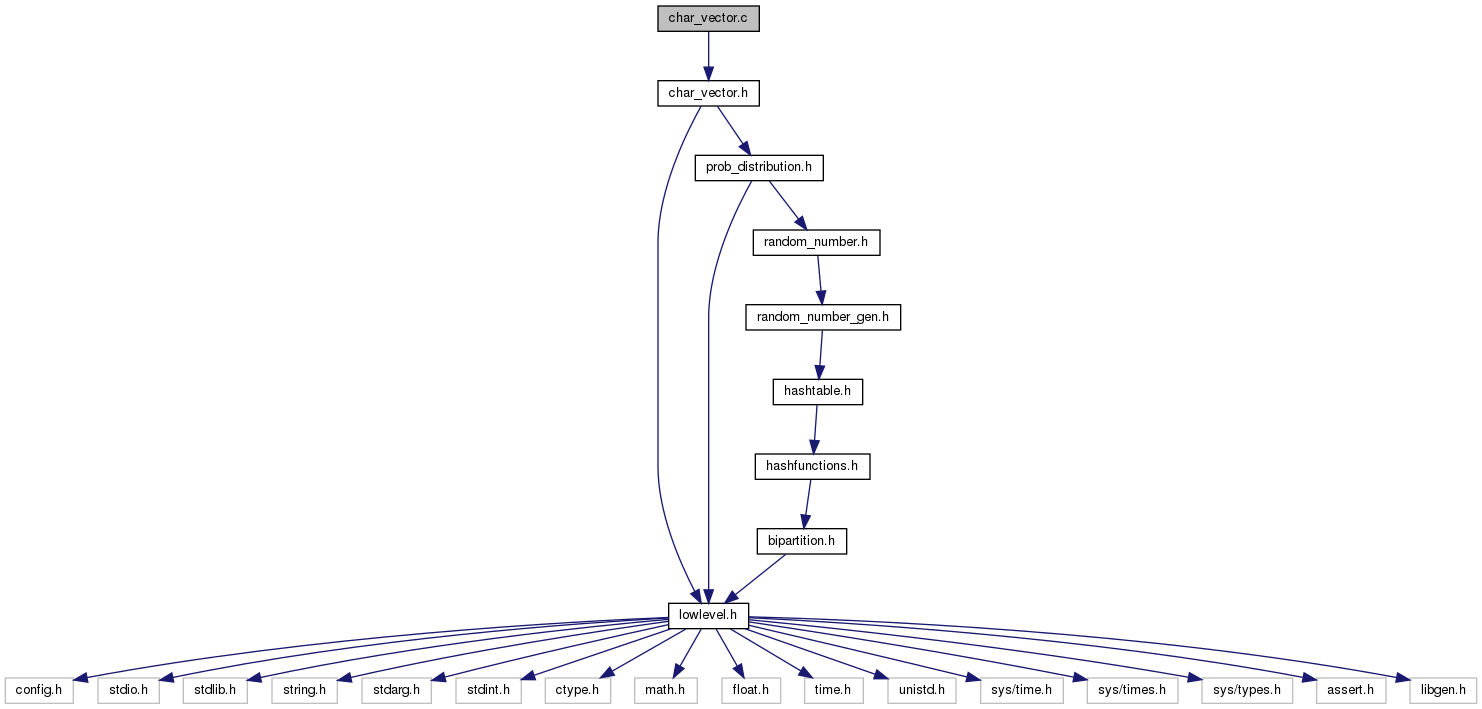
Data Structures | |
| struct | charvec_str |
Macros | |
| #define | kroundup32(x) (--(x), (x)|=(x)>>1, (x)|=(x)>>2, (x)|=(x)>>4, (x)|=(x)>>8, (x)|=(x)>>16, ++(x)) |
Functions | |
| int | compare_charvecstr_decreasing (const void *a, const void *b) |
| int | compare_charvecstr_lexicographic (const void *a, const void *b) |
| char_vector | new_char_vector (int nstrings) |
| Create a vector of strings with initial size for each string of zero. | |
| char_vector | new_char_vector_big (int nstrings) |
| Create vector of strings, and preparing it to realloc() fewer times; used in conjunction with 'append_big'. | |
| char_vector | new_char_vector_from_valid_strings_char_vector (char_vector vec, int *valid, int n_valid) |
| Create a vector of strings from subset of strings of another char_vector. | |
| char_vector | new_char_vector_fixed_length (int nstrings, int nchars) |
| Create a vector of strings where each string is assigned an initial value of nchars. | |
| void | del_char_vector (char_vector vec) |
| Delete vector of strings only after nobody is using it. | |
| void | char_vector_link_string_at_position (char_vector vec, const char *string, int position) |
| Link a previously allocated string (to avoid copying all characters) | |
| void | char_vector_add_string_at_position (char_vector vec, const char *string, int position) |
| Add a new string (vector of characters) at specific location. | |
| void | char_vector_add_string (char_vector vec, const char *string) |
| Add a new string (vector of characters) at next available location. | |
| void | char_vector_append_string_at_position (char_vector vec, const char *string, int position) |
| Append string at the end of existing string at location. | |
| void | char_vector_append_string (char_vector vec, const char *string) |
| Append string at the end of existing string at most recently used location. | |
| void | char_vector_append_string_big_at_position (char_vector vec, const char *string, int position) |
| Append strings like before, but doubling allocation space if insufficient (reduces calls to realloc() ) | |
| void | char_vector_append_string_big (char_vector vec, const char *string) |
| void | char_vector_finalise_big (char_vector vec) |
| void | char_vector_expand_nstrings (char_vector vec, int new_size) |
| Increase size of vector of strings (called automatically by other functions) | |
| void | char_vector_reorder_strings_from_external_order (char_vector vec, int *order) |
| update order of strings in vector based on a vector of new positions | |
| int | char_vector_remove_empty_strings (char_vector vec) |
| Reduce size of vector of strings by removing empty strings (returns number of empty strings) | |
| int | char_vector_remove_duplicate_strings (char_vector vec) |
| Remove identical strings and resizes char_vector_struct. | |
| void | char_vector_reduce_to_valid_strings (char_vector vec, int *valid, int n_valid) |
| reduce char_string_struct to only those elements indexed by valid[] | |
| void | char_vector_reorder_by_size_or_lexicographically (char_vector vec, bool lexico, int *order) |
| Order char_vector_struct elements from longer string to smaller, or lexicographically; can be used after calling new_char_vector_from_file() but if topology etc. are associated to it, then order[] must be externally defined and will have new locations, to keep track of changes. | |
| bool | char_vector_link_address_if_identical (char_vector *v1, char_vector *v2) |
| If the two char_vectors are identical (same strings in same order), then delete one and make it point to the other one. | |
| void | index_species_gene_char_vectors (char_vector species, char_vector gene, int *sp_idx_in_gene, int *order_external) |
| find occurences of species->string[] inside gene->string[] filling indexes in sp_idx_in_gene. More... | |
| void | update_species_count_from_gene_char_vector (char_vector species, char_vector gene, int *sp_count) |
vector of strings (species names, leaf names, etc.)
| void index_species_gene_char_vectors | ( | char_vector | species, |
| char_vector | gene, | ||
| int * | sp_idx_in_gene, | ||
| int * | order_external | ||
| ) |
find occurences of species->string[] inside gene->string[] filling indexes in sp_idx_in_gene.
The species are taxon names which may be associated with topologies or alignments, such that we can not reorder its elements here (without also modifing e.g. tree leaves). But ordering from longer to shorter is essential for pattern finding, so it is assumed that the char_vector is already sorted UNLESS user provides the ordering.
 1.8.13
1.8.13