SpecImage module for python
A general module for hyperspectral image analysis
Developed by Leonardo de Oliveira Martins at the University of Vigo and at the Imperial College London for the SAVVY Project.
Main Features
- Import text files from WiTec and Matlab; saving to Pickle format
- Plotting of spectral densities
- Removal of noise and cosmic spikes from spectra
- Interpolation and rescaling (normalization)
- Image simplification/reduction
- Background and baseline corrections
- Endmember extraction and abundance maps
- Modelling, clustering and classification
Noise and cosmic spikes
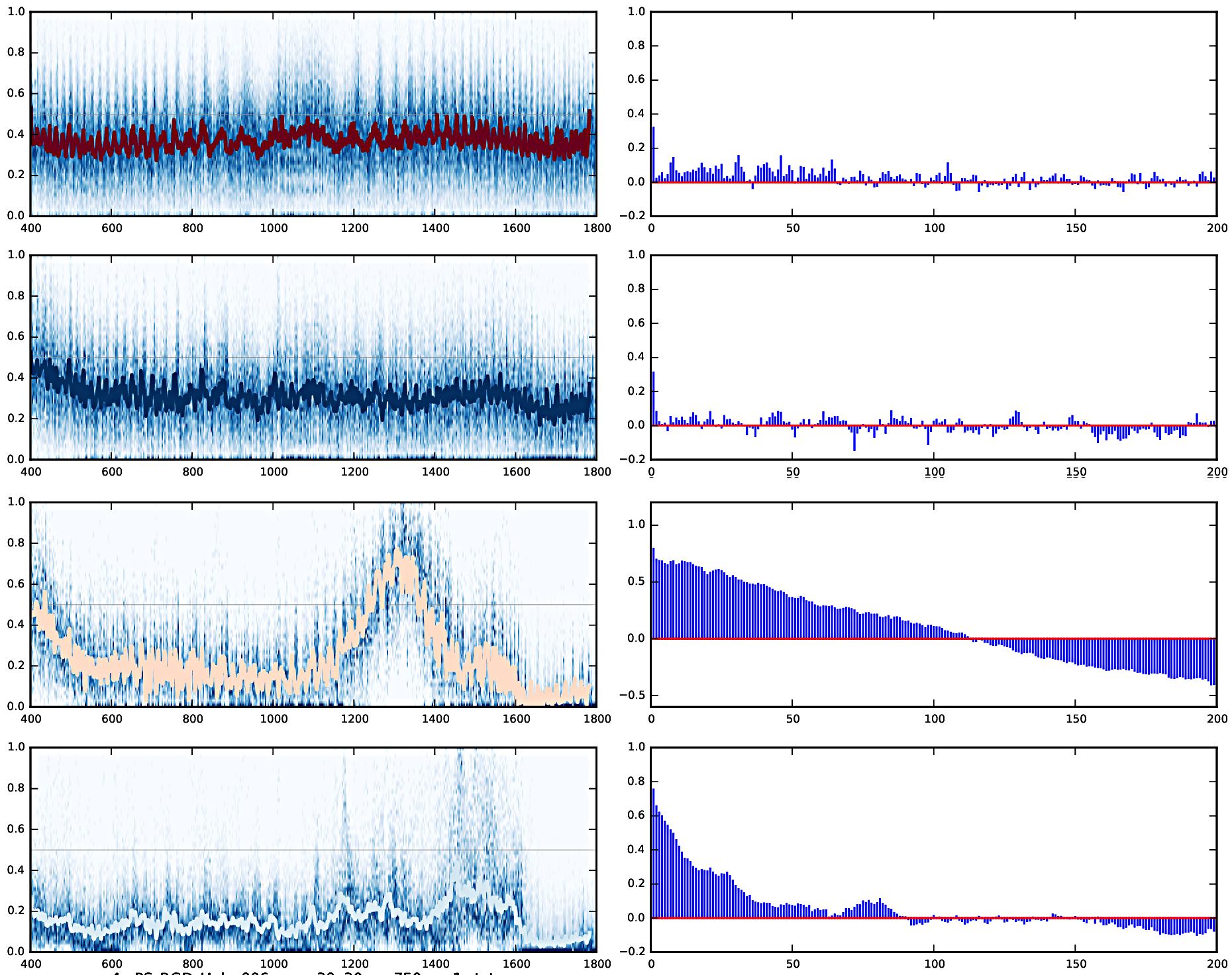
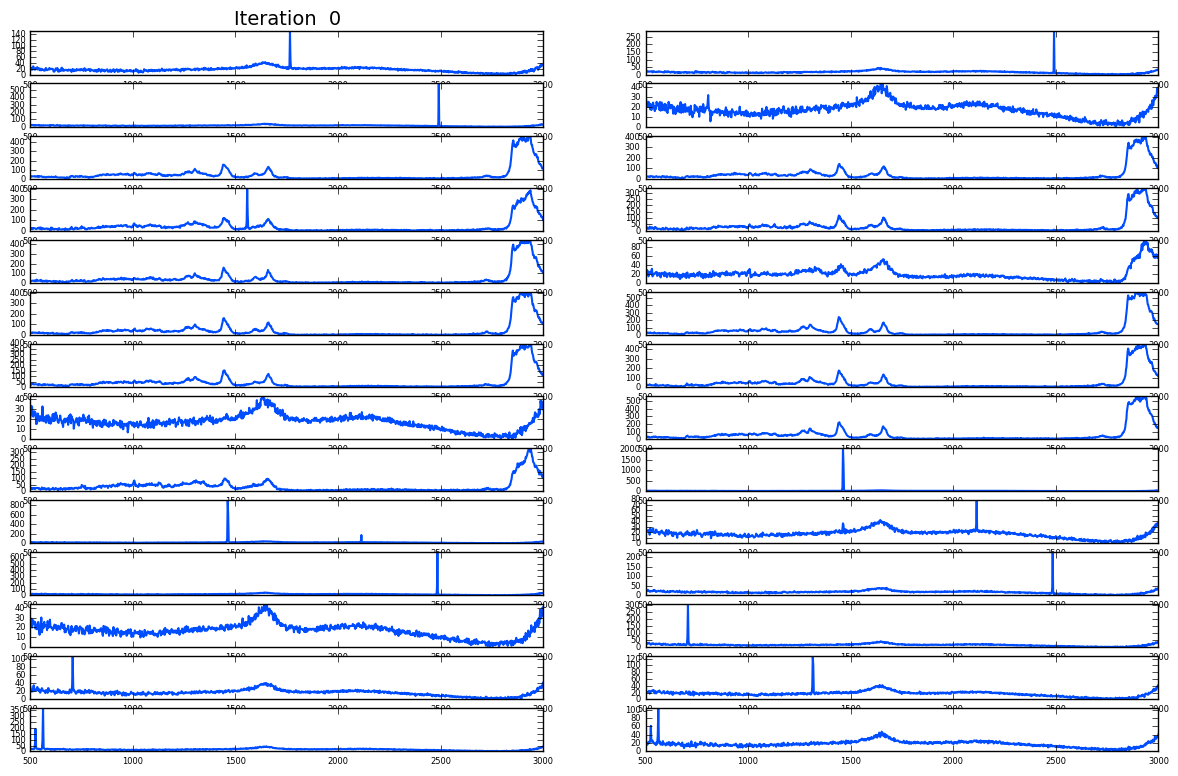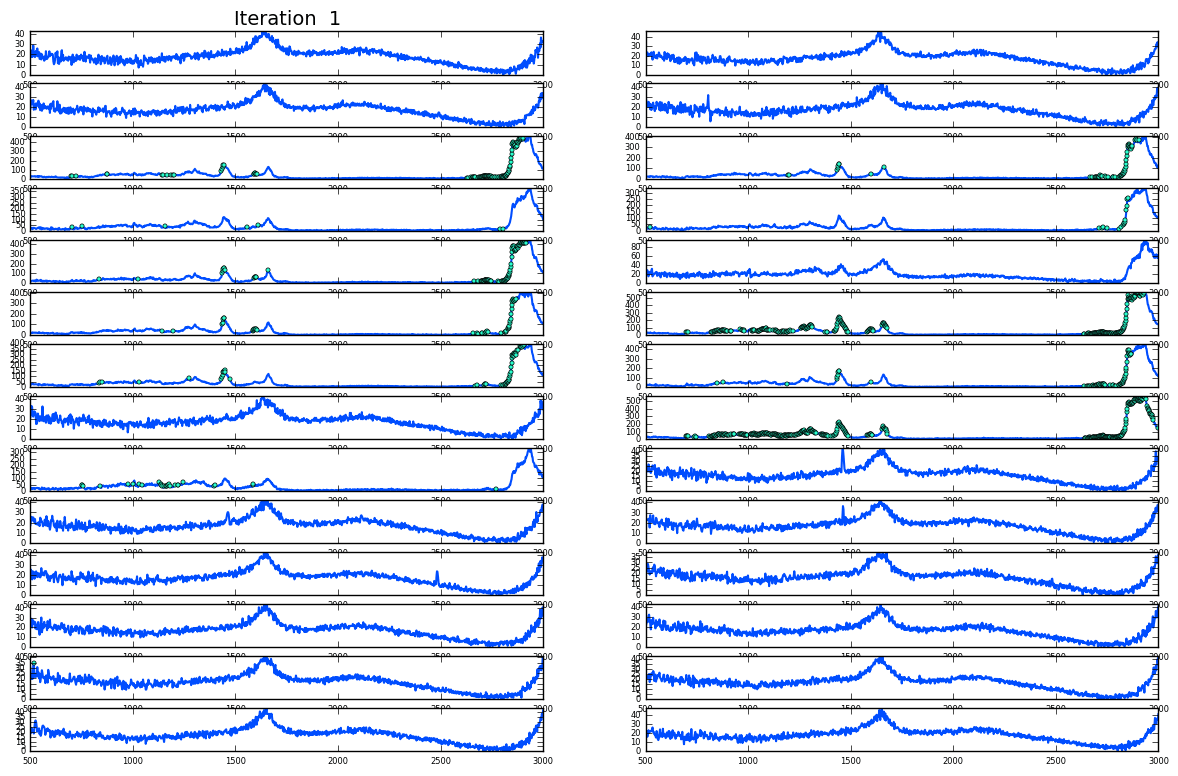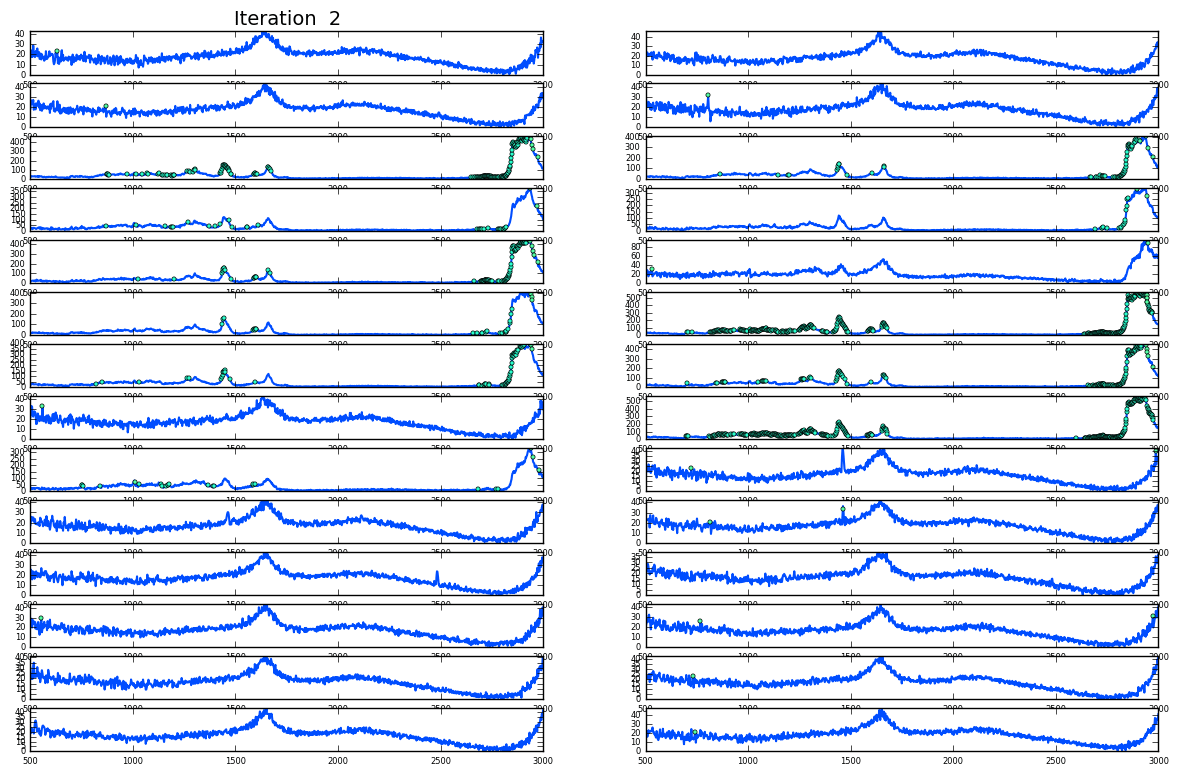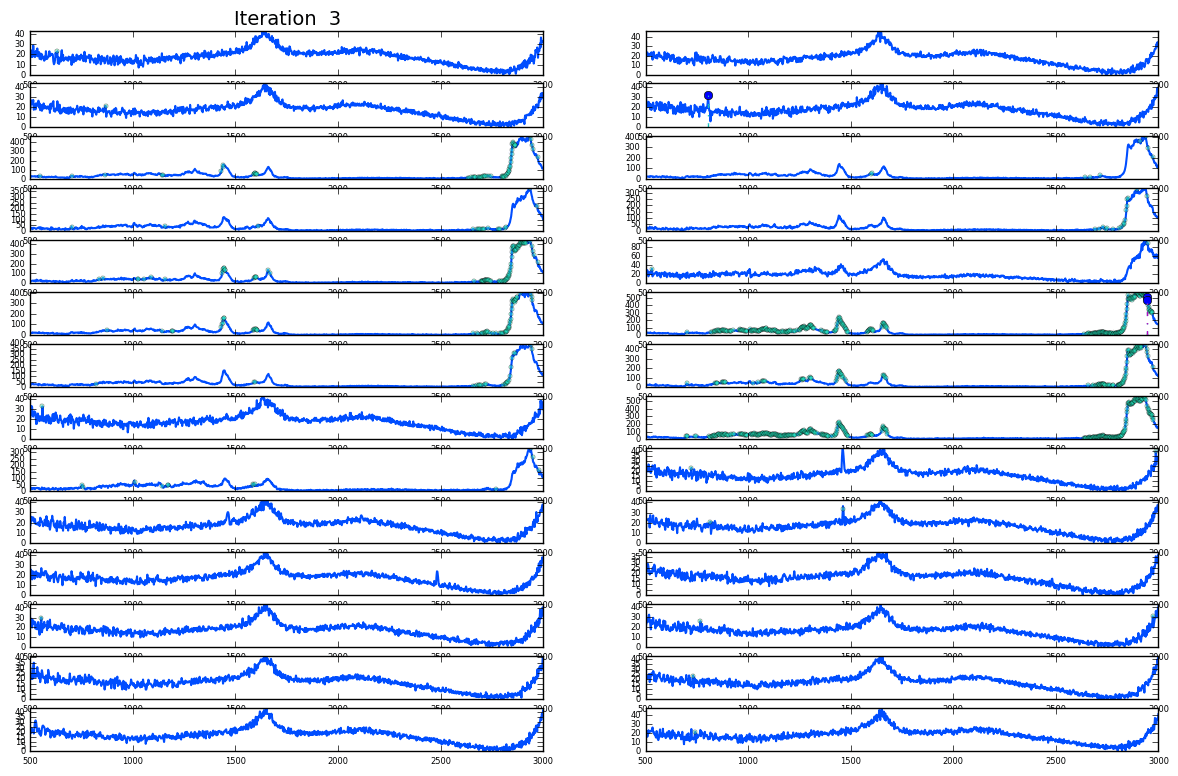
Plotting of distribution of spectra

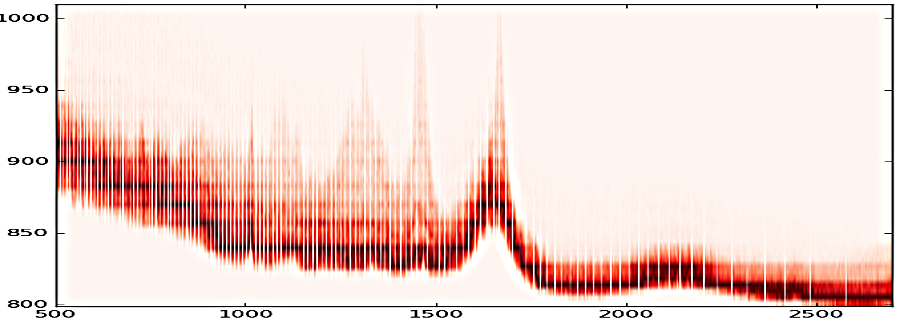 OBS: the images represent distinct images
OBS: the images represent distinct images
Interpolation
Savitzky–Golay filter
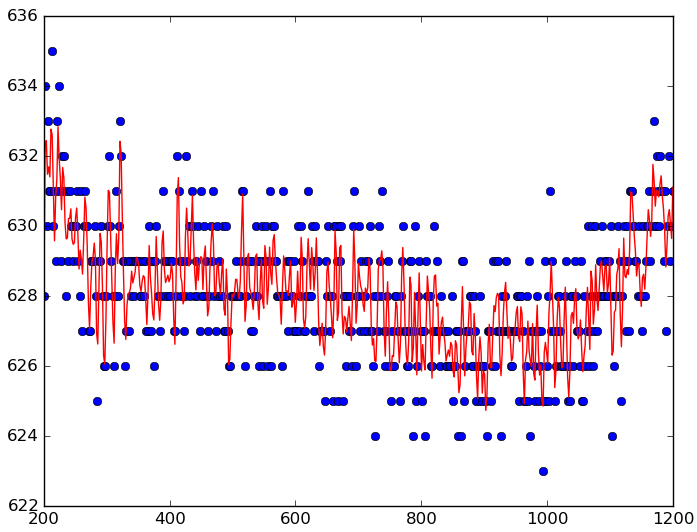
|
Moving average smoothing
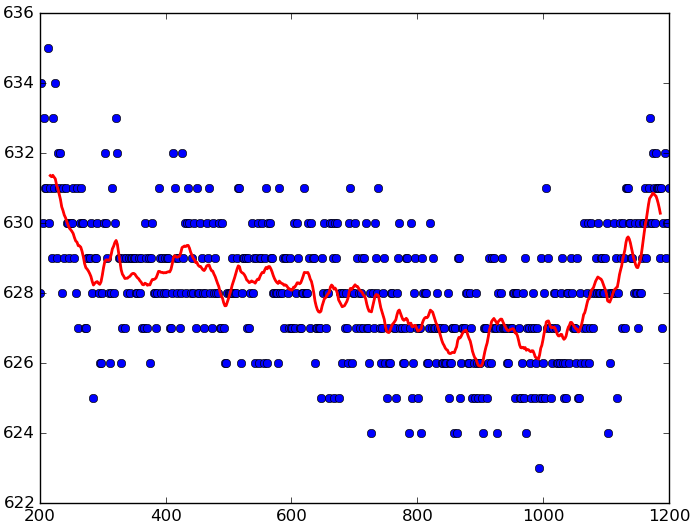
|
Also
- Linear interpolation
- Spline interpolation
- LOWESS regression
Prediction via interpolation
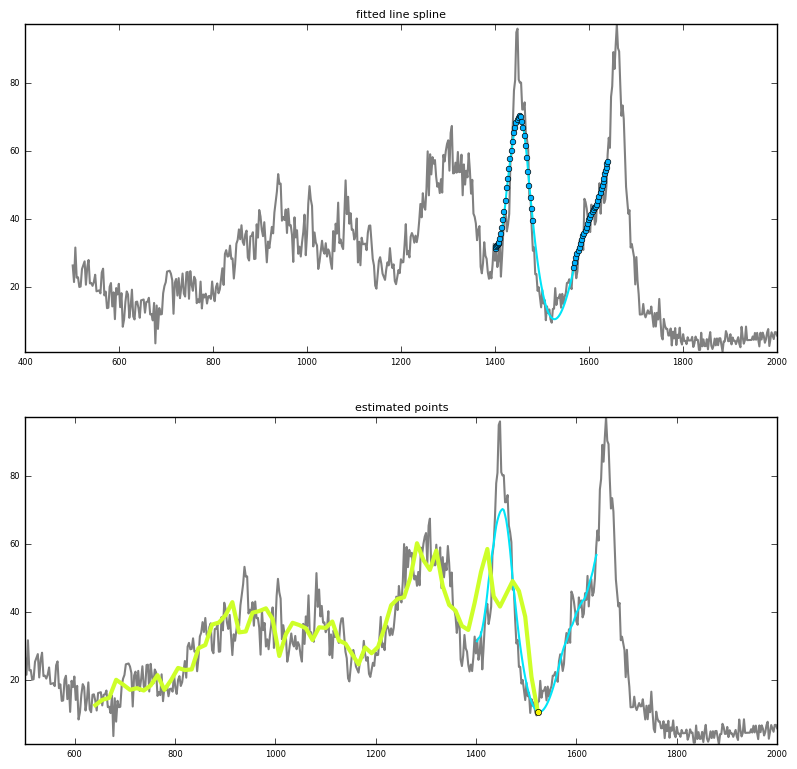
|
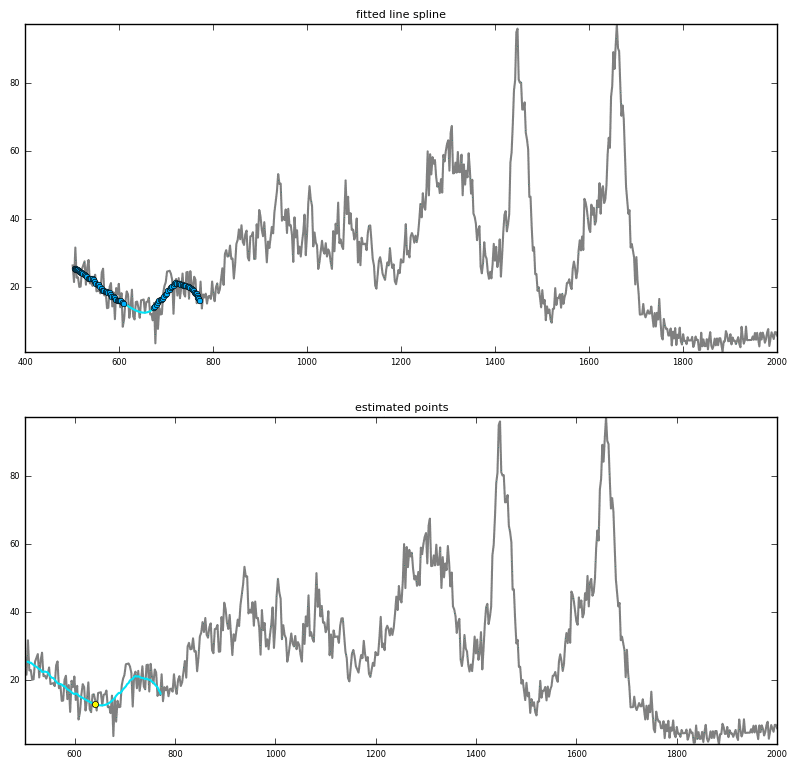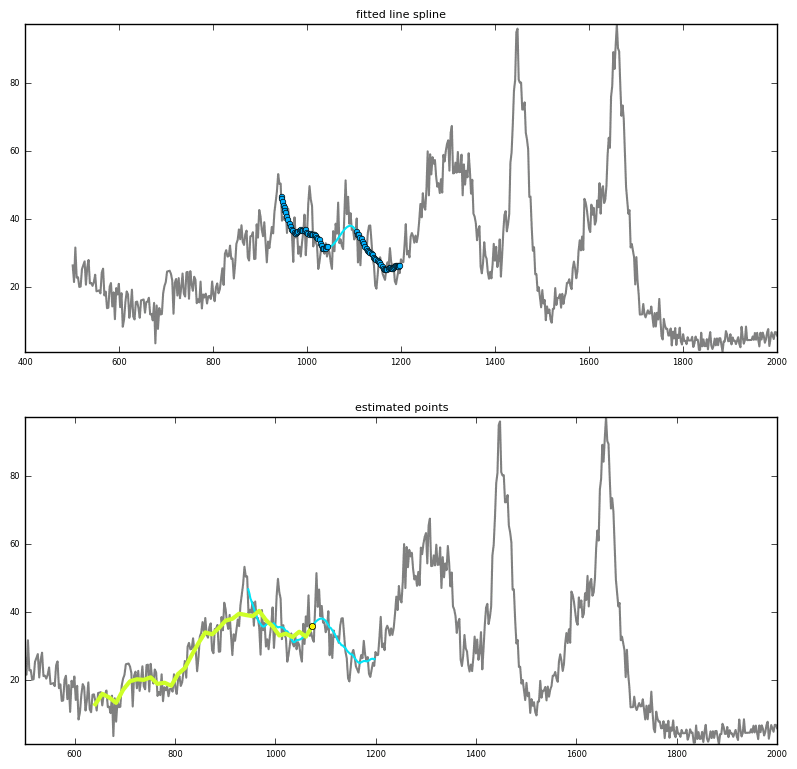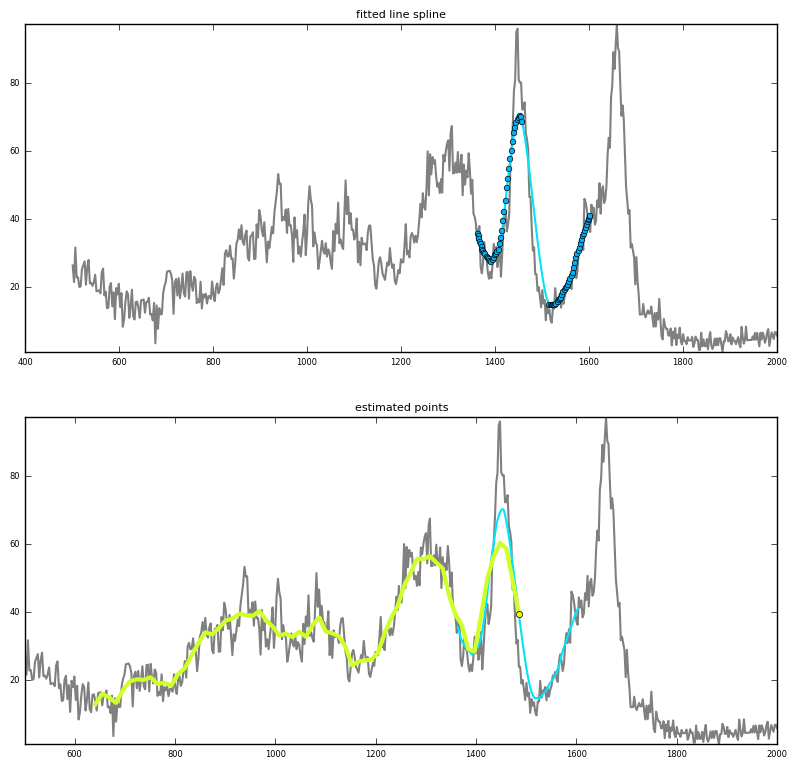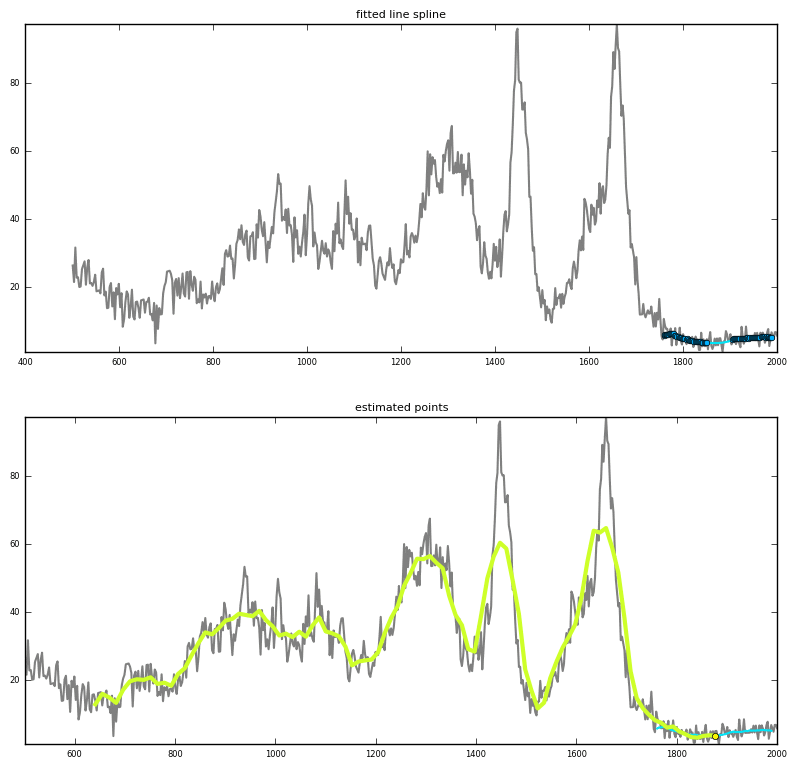
|
Normalization
def rescale_01(self):
# Linear rescaling to interval zero-one
self.spc = np.array([rescale_spectrum_01(spec) for spec in self.spc],dtype=float)
return self
def rescale_zero(self):
# force series to start at zero, while maintaining range
self.spc = np.array([spec - min(spec) for spec in self.spc],dtype=float)
return self
def rescale_mean(self):
# centralization to the mean
self.spc = np.array([spec/np.mean(spec) for spec in self.spc],dtype=float)
return self
def rescale_zscore(self):
# Standardization of spectrum to a N(0,1)
self.spc = np.array([(spec - np.mean(spec))/np.std(spec) for spec in self.spc],dtype=float)
return self
Simplification of spectral matrix
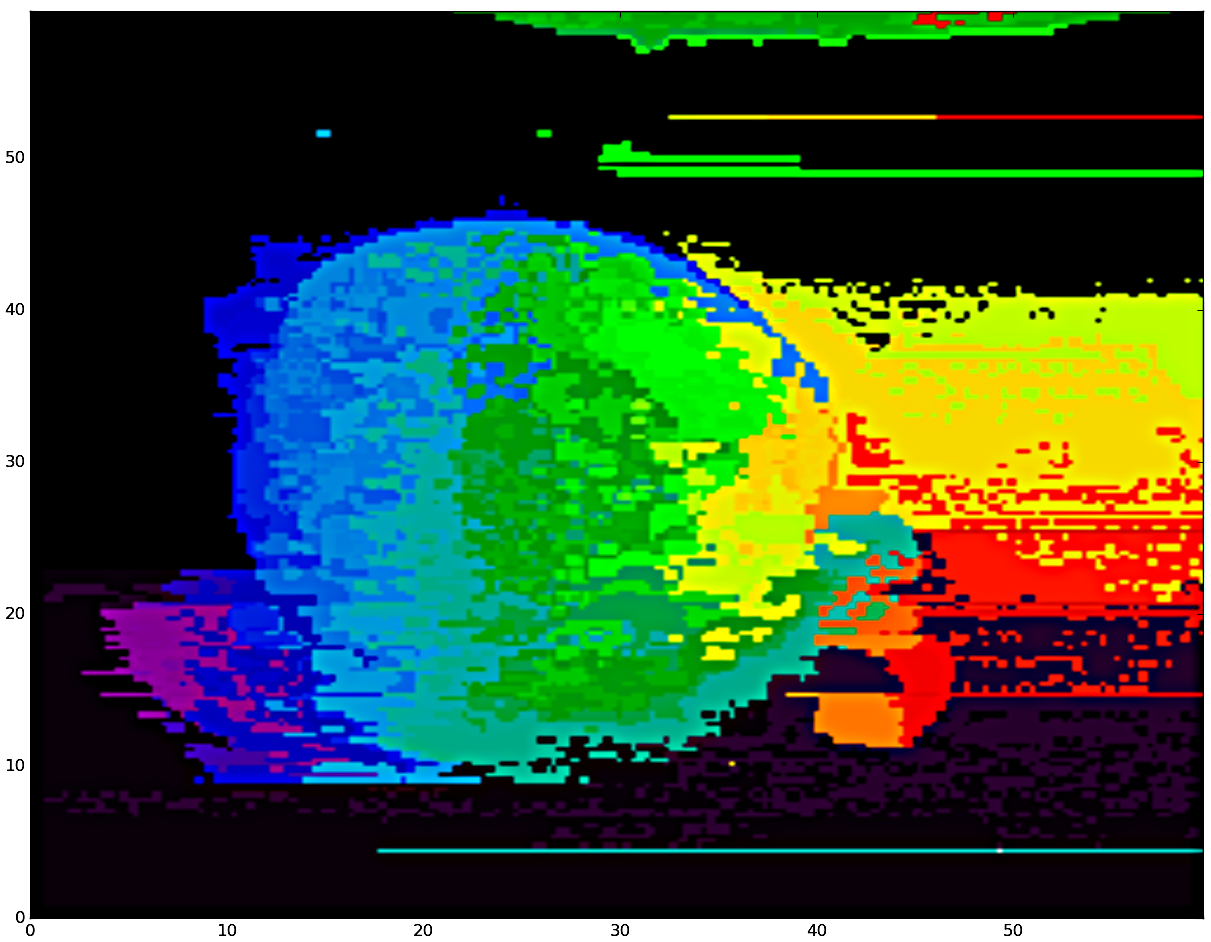 Similar pixels replaced by 'exemplar' spectra
Similar pixels replaced by 'exemplar' spectra
|
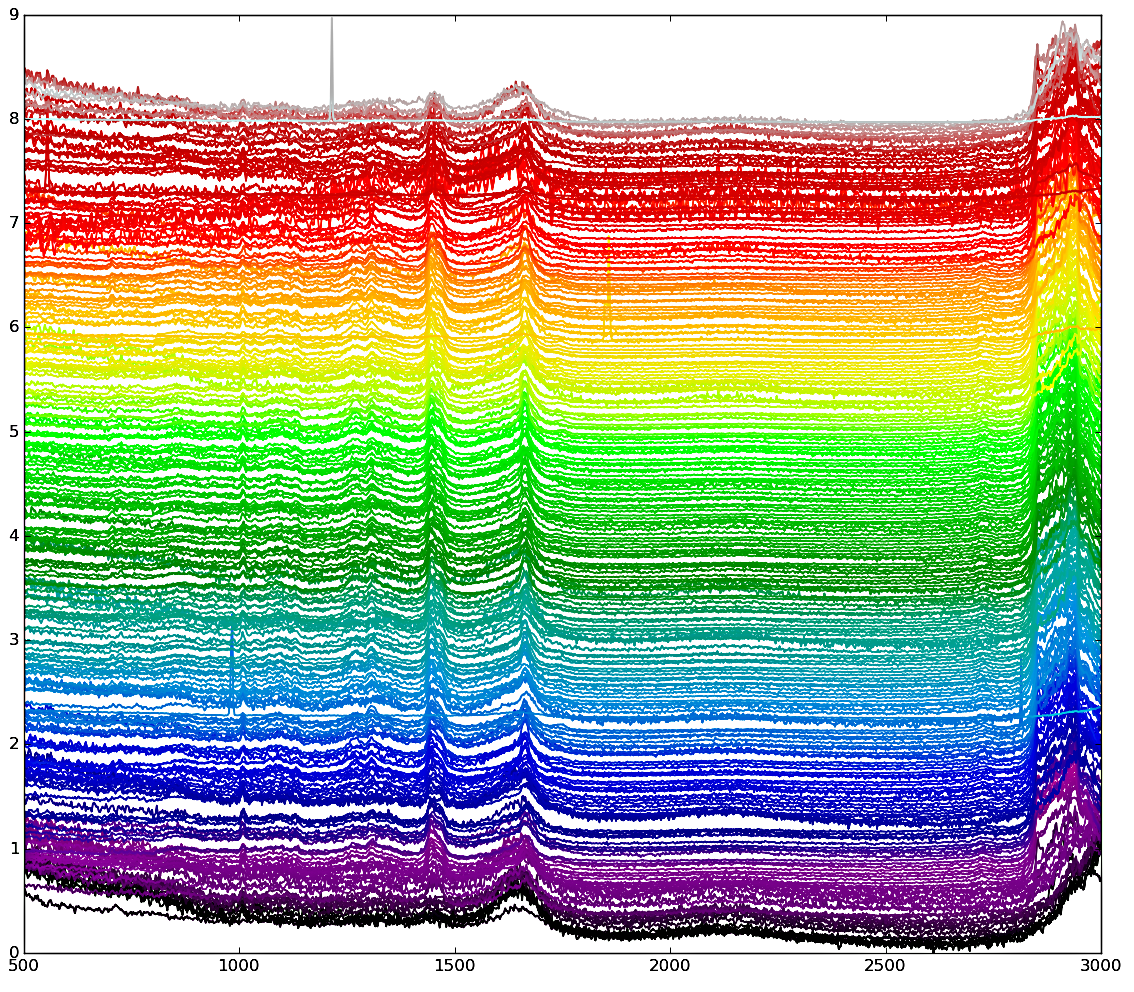 List of 'exemplars'
List of 'exemplars'
|
Background correction by PCA transform
- Application of a PCA dimensionality reduction followed by inverse transform
- Recreates original data but using only principal components
- Can be applied over wave numbers or over spectra (pixels)
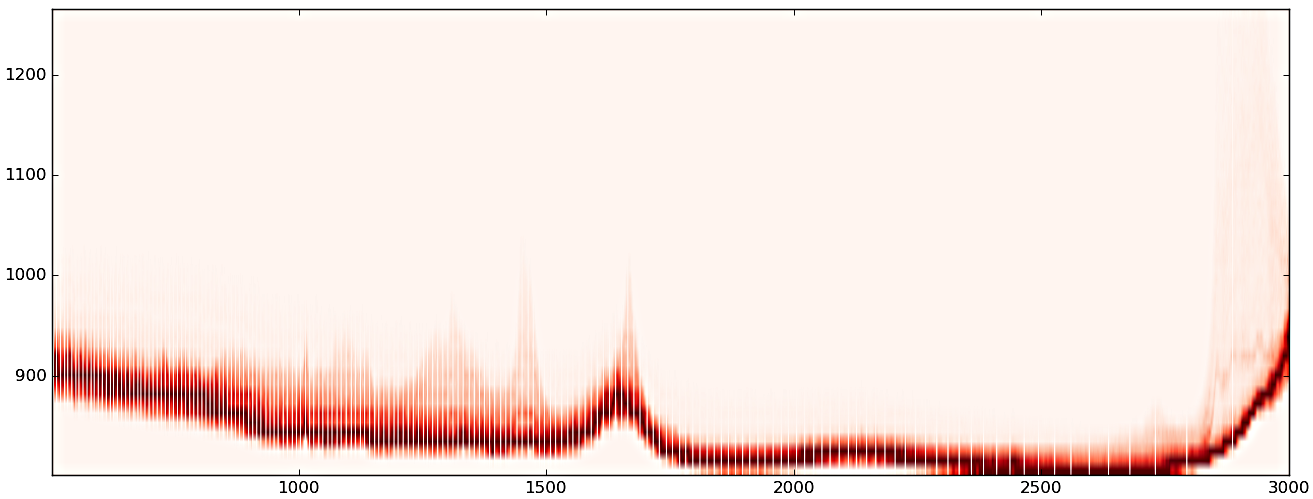 (1) Original data
(1) Original data
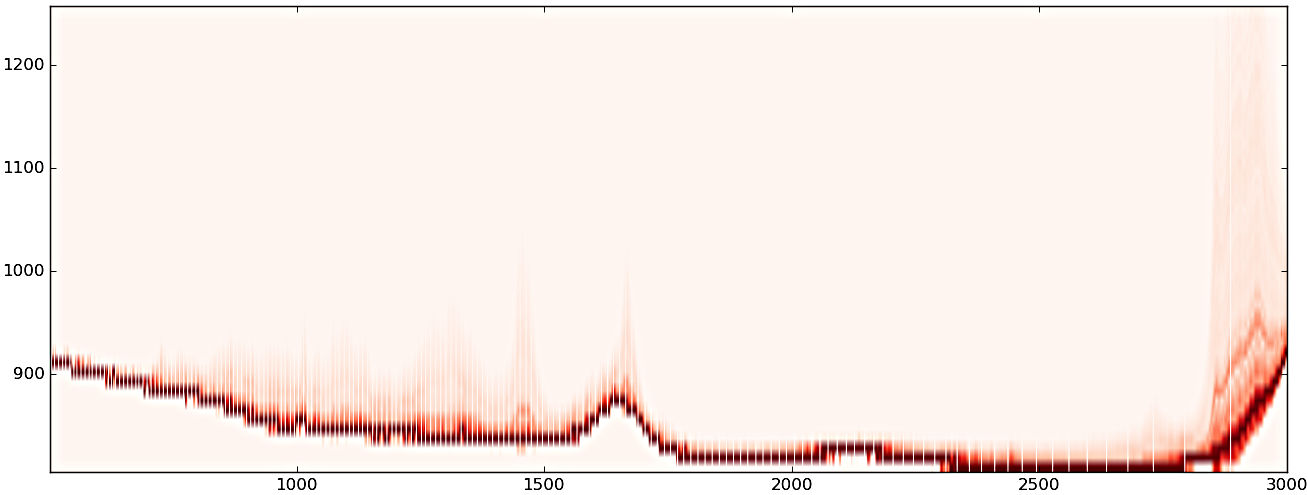 (2) after PCA noise reduction
(2) after PCA noise reduction
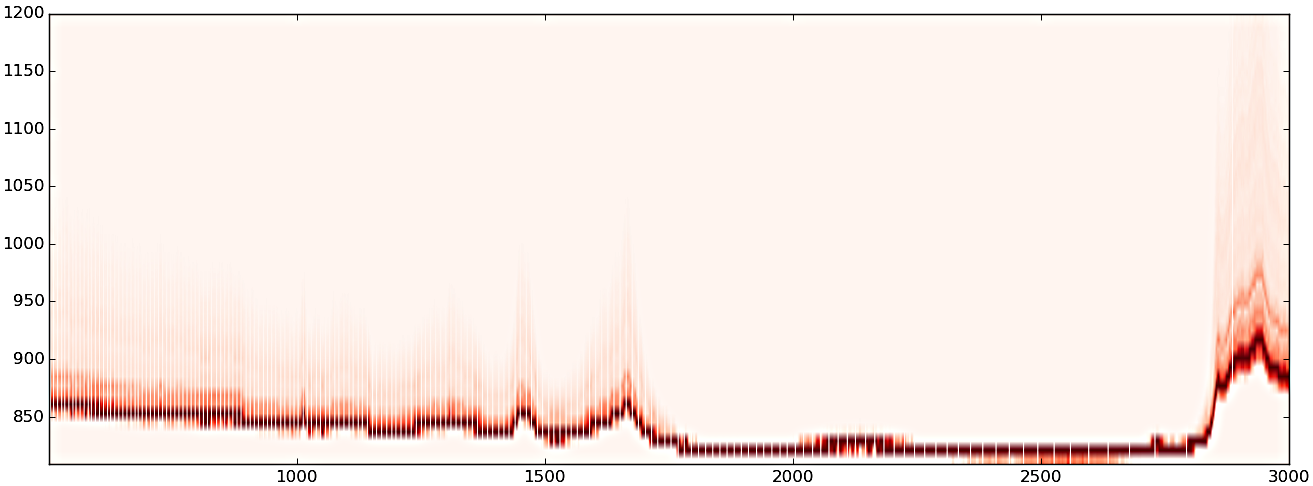 (3) after PCA pixel reduction
(3) after PCA pixel reduction
Baseline correction
Rubberband baseline
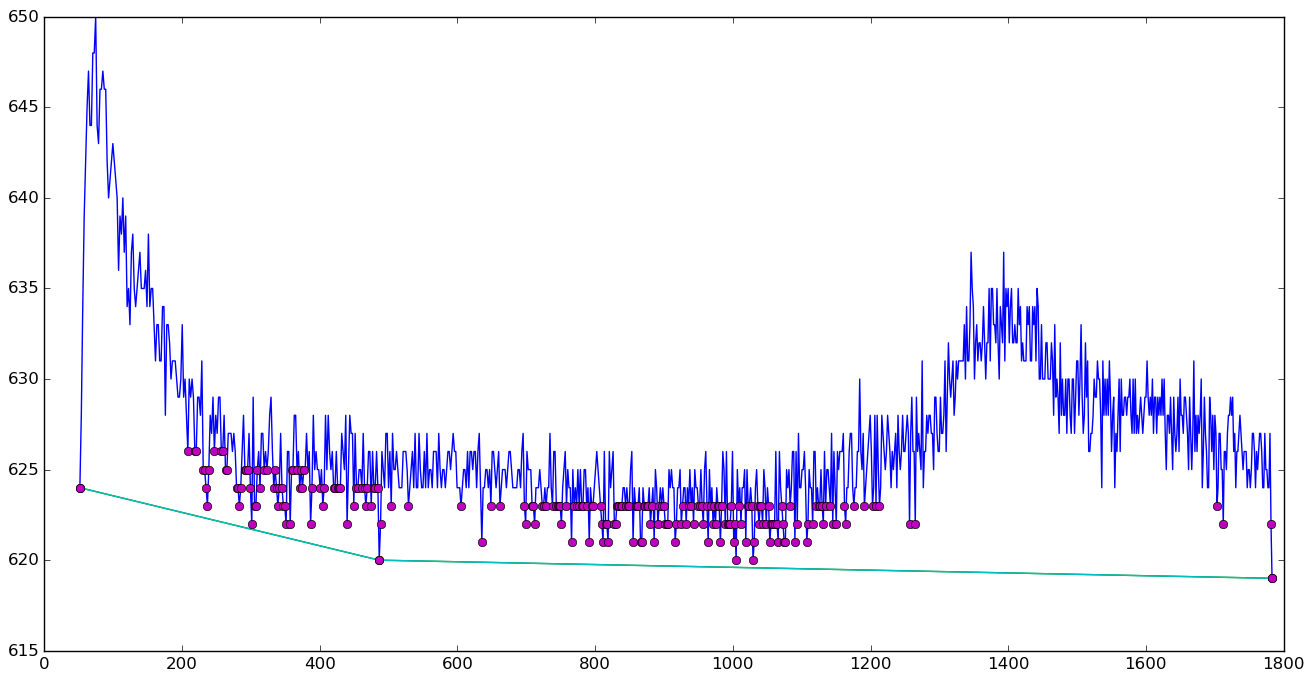
|
Polynomial baseline
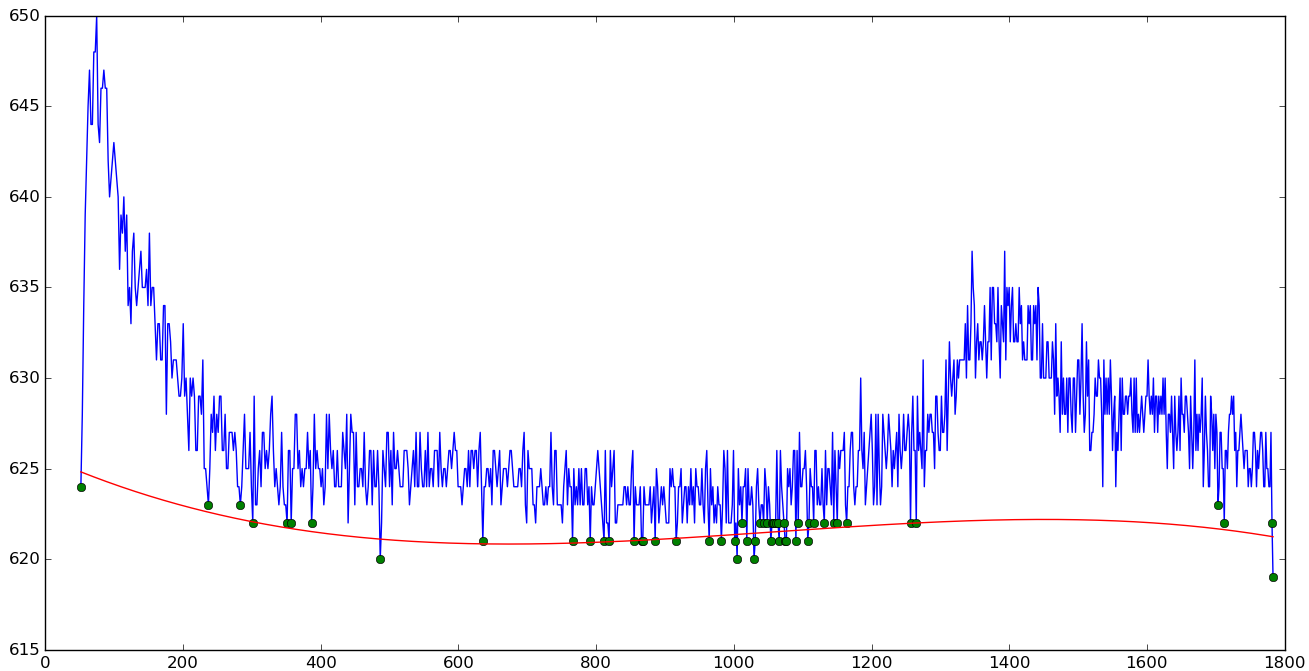
|
Piecewise rubberband baseline
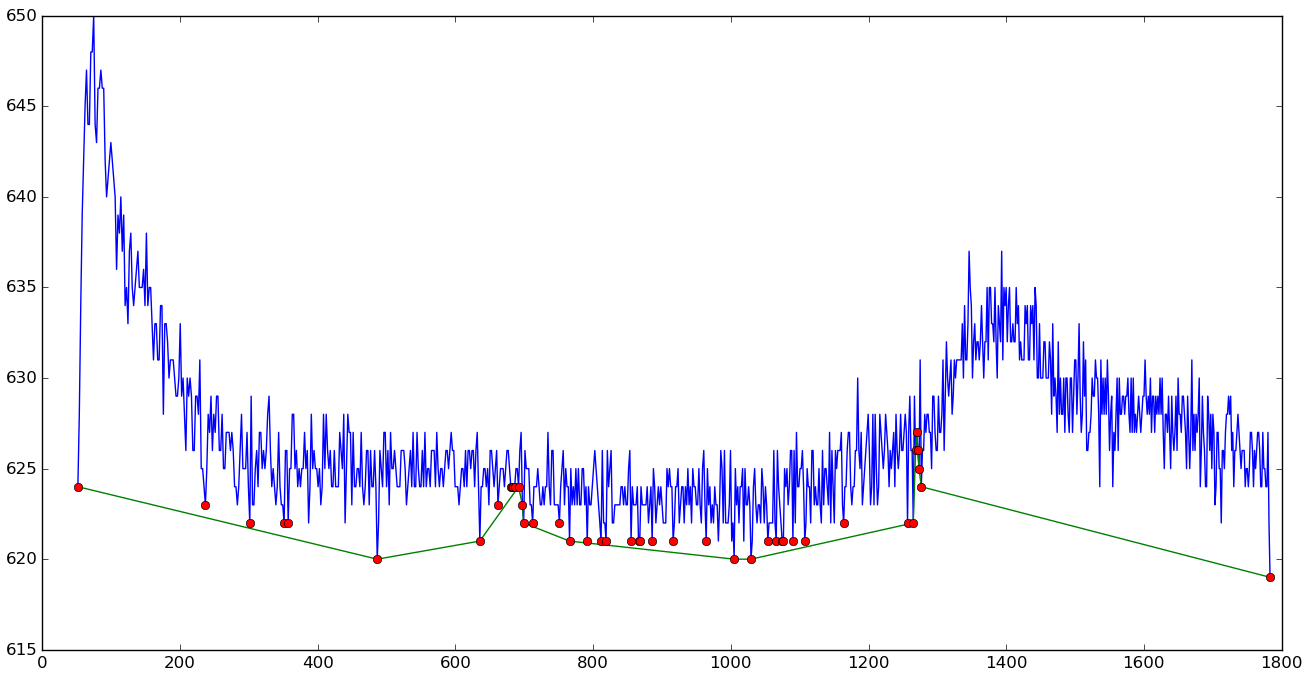
|
Asymmetric least squares baseline
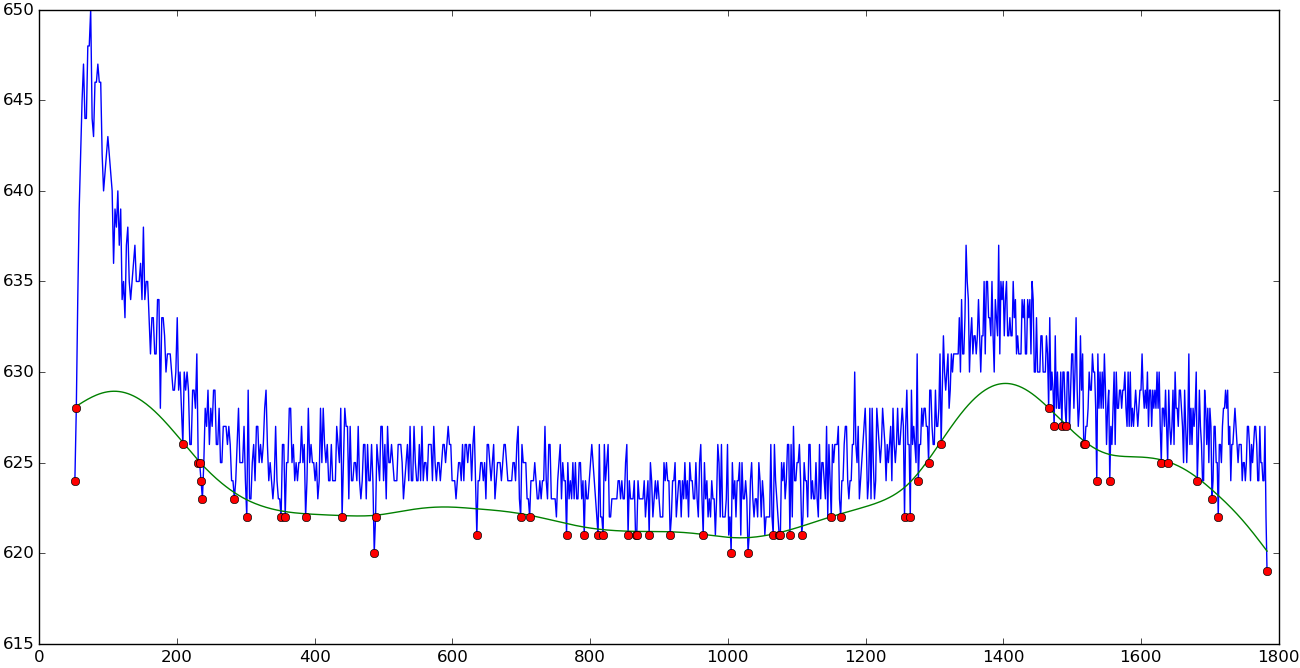
|